
|

|

News
About TARIS TARIS (Tree Analysis and Representation of Isopotential Surfaces) is a software package designed for molecular similarity purposes in the field of Computational Chemistry. Users are allowed to compare molecules through their molecular electrostatic potentials (MEP) loading .cube files according to the GAUSSIAN format. TARIS performs a geometrical and topological analysis of the MEP for each molecule going through a potential scan. In addition to the .cube file, the user must provide the minimum and maximum cutoffs as well as the stepsize for the potential scanning. This information is encoded in a rooted tree where the nodes are weighted by positive real numbers. In order to compute a dissimilarity measure among molecules, a tree edit distance measure has been implemented. TARIS output returns the distance matrix, the maximum distance in the whole molecular set and the similarity matrix (derived from the distance matrix). The whole code is written in C++ and has been tested in gcc and intel compilers and it works in any UNIX based platform. 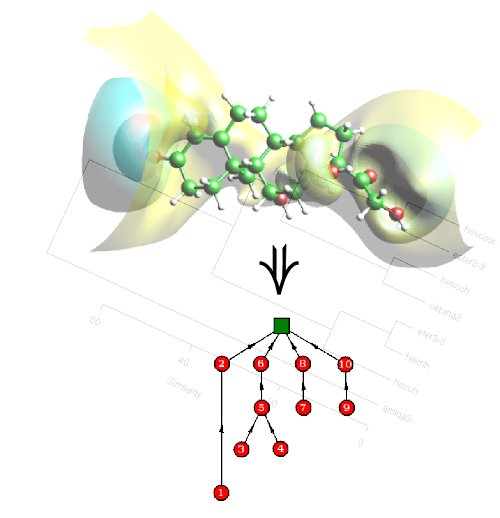 Authors
|
